
The cosine similarity always belongs to the interval. This will compute the cosine similarity between the test documents and the training documents. The cosinesimilarities variable will be a 2D array with shape (ntestdocuments, ntraindocuments). It follows that the cosine similarity does not depend on the magnitudes of the vectors, but only on their angle. This will compute the cosine similarity between the test documents and the training documents. Cosine similarity is the cosine of the angle between the vectors that is, it is the dot product of the vectors divided by the product of their lengths. 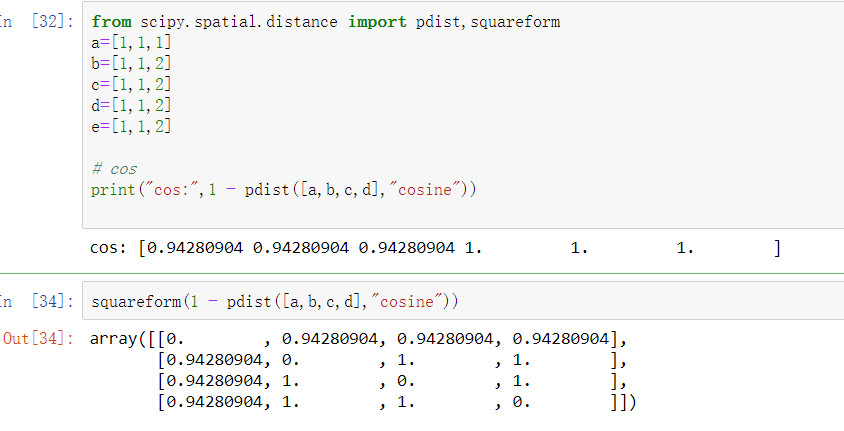
I want to know that am i doing it right ? Is it right to calculate the cosine similarity first and then pass the values to kmeans ? As by default, the sklearn's kmeans function uses euclidean distance.In data analysis, cosine similarity is a measure of similarity between two non-zero vectors defined in an inner product space. I am also getting a list of clusters for the users. The characteristic array of the sample spectrum was compared with that of standard spectra to obtain the cosine similarity score using the Sklearn library. On L2-normalized data, this function is equivalent to linearkernel. 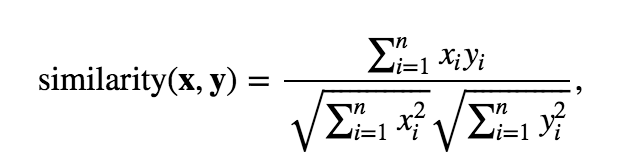
With this code, i am getting the cosine similarity for all the pairs and i can filter the pairs based on the value of cosine similarity. Compute cosine similarity between samples in X and Y. Do you think that scaling all vectors to be on the unit sphere and then run the sklearn Kmeans is identical to running k-means with cosine similarity. from import cosinesimilarity import numpy as np X np. when I used Spark to to so, I had no problem. Kmeans = KMeans(n_clusters=2, init ='k-means++', max_iter=50, I have embedded vectors with K-means using cosine similarity. Pairs = pairs.to_frame('cosine distance').reset_index() cosinesimilarity accepts scipy.sparse matrices. Here will also import NumPy module for array creation. The code is: import pandas as pdįrom import cosine_similarityĭf = pd.read_csv('input_file.csv', sep=',', encoding='latin-1', This kernel is a popular choice for computing the similarity of documents represented as tf-idf vectors. Step 1: Importing package Firstly, In this step, We will import cosinesimilarity module from package. I measured cosine similarity and then clustered the users with sklearn's cosine_similarity and kmeans clustering algorithm. The rows are users and M in the column is movie and C is the choice. M200Īccording to the scheme, if an original review was 1 (good) then we put 1 in the cell and -1 in the cell if the review was 0 (bad). Addition Following the same steps, you can solve for cosine similarity between vectors A and C, which should yield 0.740. It returns a matrix instead of a single value 0.8660254. The csv file is shown below: UserID M1 M2 M3. I am using below code to compute cosine similarity between the 2 vectors. I used cosine similarity measure with k-means clustering. I want to cluster similar users based on their reviews with the idea that users who rated similar movies as good might also rate a movie as good which was not rated by any user in the same cluster. A value of 1 for good, 0 for bad and blank if no choice. I have a movie dataset with more than 200 movies and more than 100 users.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
